Work in Denmark has laid the foundations for identifying all the microbes in wastewater treatment systems and anaerobic digesters around the world with the aid of genetic fingerprinting. By Per Halkjær Nielsen.
Microorganisms are the workhorses of wastewater treatment systems and anaerobic digesters, where they are responsible for removal of pollutants and pathogens, recovery of nutrients and energy, and producing clean water.
Numerous different microbes exist in these systems; however, just how many is unknown, with information about their identity and aspects of function available only for a few. Without such knowledge, it is difficult to make informed decisions on management and optimisation of the treatment systems. This will be especially important in the years to come, given there is a global trend towards moving from simple wastewater treatment to advanced water resource recovery systems for sustainable development.
What can instagram teach you about natural bodybuilding rhyder promotions sustanon 250 gains mechanical tension – dr. giunta federico – sports nutritionist rome – fitness and bodybuilding.
Novel DNA-based technologies have radically changed this landscape. It is now possible to identify microbes to species level, and this has been done for the first time for all microbes in Danish wastewater treatment plants and digesters. In addition, mapping of all microbes in these systems worldwide is ongoing.

of bacteria by molecular
methods. The
fingerprint gene, 16S
rRNA gene, is present
in all bacteria and its
DNA sequence can be
used for identification
by comparing it to a
reference database (if it
is present in this)
The gold standard for identification of microbes, without the need to first grow them in a pure culture, is to identify special marker genes in their chromosome – a fingerprint gene (Figure 1). This can be achieved directly from a sample of activated sludge or digester sludge. The DNA sequence of this marker gene is then checked against a reference database and, if the gene is present, the bacteria can be identified. However, the big problem is that existing databases are underpopulated and only contain a limited number of sequences, especially the relevant ones that come from sludge, so the majority of microbes from wastewater systems remain unidentified.
The new method
Our team has developed a new sequencing method that has addressed these shortcomings. We have now made a new ecosystem-specific database that contains sequences from nearly all microbes present in Danish wastewater treatment plants and digesters, called MiDAS (Microbial Database for Activated Sludge). Since most of the microorganisms are new and undescribed, we have also developed a new MiDAS taxonomy – an identification system that allows all species to be given a unique ID, so they can be recognised in future studies in any part of the world. We know that many of the microbes in Danish plants are also abundant across the world, so use of the new identification systems (the reference database and taxonomy) is useful for all working in the field. Furthermore, we are currently working on a global survey that will enable us to make a near-complete reference database for microbes found in wastewater treatment plants and digesters worldwide. We expect this to be available in early 2020.
“Another inspiration for us to gain understanding is the plant operators”
Such a comprehensive database is really needed. So far, it has been impossible to identify the microbes to species level, and it is also often difficult to achieve identification at a genus level because of limitations of existing public databases. With species resolution, we can now start comparing results from various world studies, and begin linking microbes’ identity to their function. Eventually, we will be able to link the function and importance for operation to all relevant microbes. We have already started to collect all available information, which can be viewed on the project website: MiDAS Field Guide (www.midasfieldguide.org). With this tool, we aim for a collective effort and hope other researchers in the field will help to complement the guide by providing new information when it becomes available.
The Danish plant survey
One of the key questions in microbial ecology is, how many microbial species exist in a given ecosystem. The survey of Danish plants has shown that in a typical treatment plant 2–3000 species are present. However, most are in very low abundance, and presumably are not important to the treatment processes, with only a few hundred being abundant and important. There exists a significant similarity in microbial community structure among most plants, especially where they have the same overall process design (e.g., nitrogen removal), so the total number of species across all Danish plants is 3–4000. Our best estimate for the total number of species across the world is 25–30,000, and we look forward to finishing the global survey and answering that question.

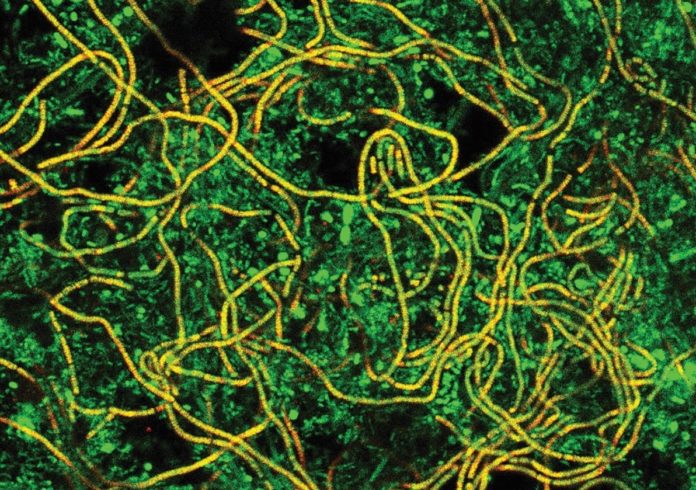
Most of the species found are novel and unknown, and many also belong to undescribed genera. This underlines the huge undiscovered diversity in these systems, which is sometimes called “microbial dark matter”. It is indeed a challenge to reveal the functions of those microbes, but we have a range of approaches to achieve that. When we know their fingerprint gene, we can also design markers to visualise them by fluorescence microscopy, a technique called fluorescence in situ hybridisation (FISH). Microbes in the wastewater treatment plants can have many morphologies but are often small rods growing in small colonies. Some form long filaments, sometimes millimetres long (see Figure 2a), and they can cause serious operational problems, such as poor settling in the clarifier or foaming. We can combine FISH with other stainings or spectroscopic techniques, such as Raman microspectroscopy, and in this way obtain information about physiological traits – for example, whether they are polyphosphate-accumulating organisms (Figures 2b and 3), carry out nitrification or consume specific nutrients.

storage polymers in Ca.
Accumulibacter and
other polyphosphate
accumulating
organisms with
fluorescence in
situ hybridisation
and Raman
microspectroscopy
The future for novel developments
Novel developments in sequencing technologies are now also paving the way for gaining access to the entire genomes of most key organisms in activated sludge systems and digesters. We will soon be ready with more than 1000 high-quality genomes from the Danish wastewater treatment plants, serving as a blueprint of important information about their physiological potential and function in wastewater treatment systems. They can be linked to the fingerprint genes in the MiDAS reference database, providing an invaluable resource.
One of the ultimate goals for operators at wastewater treatment plants is to enable online identification of the microbes and use that to monitor and control the microbial communities (Figure 3). This will soon be possible. With the newest sequencing technologies, it is possible to bring pocket-sized sequencing equipment to the treatment plants and get the results in a few hours, making this fast and cheap. With the new MiDAS reference database, fast and reliable identification is now possible. The result could be that troublesome filamentous bacteria can be identified and advice about control measures given; it may be possible to evaluate whether the right amount and composition of nitrifiers is present; or to test the effluent quality, for example, in relation to the presence of pathogens. There are numerous options, and many we have not thought of yet.
The MiDAS project experience

of ‘online’ monitoring
and potential control of
microbial communities
With all the work described here, we are gaining a lot of experience. More than 50 wastewater treatment plants have been part of the MiDAS project since 2006. From this, we have collected extensive and comprehensive datasets providing community composition information for different full-scale plants, primarily with nutrient removal, including variations over the years. Most bacterial species are present in all plants but in varying abundances. We do not always understand why, but the new reference database and online identification will speed up deciphering factors driving the communities.
Another inspiration for us to gain understanding is the plant operators. As soon as we get sequencing results from their treatment plant, we make it available online so plant operators in their offices or via smartphone can follow the community composition of their plant. Currently, such data are delayed some months because of the time needed for conventional sequencing but, in the future, when it comes online, the results will be delivered daily or weekly, and will be extremely useful for plant surveillance and control. The operators can also report to consultants or researchers when they experience “interesting” cases.
The microbes in anaerobic digesters at wastewater treatment plants are also identified and included in the reference database. Many species are only present because they are added with the feed, such as primary sludge or surplus activated sludge, and are thus not active. The active key anaerobic microbes are, however, less characterised compared with activated sludge microbes. The new MiDAS reference database has allowed us to reveal species distribution and key operational factors determining their presence during a six-year survey of more than 40 Danish digesters. Apart from surveillance, it is still difficult to achieve informed management of digester communities, but the next few years of experience will change this.
Conclusion
The ability to identify important microbes in the environment is now available, and we can begin to apply this new knowledge to solving many problems, including challenges that have not yet been considered. The new sequencing method and ecosystem-specific reference database will enable sharing of the information from all relevant world studies, so that when a problem occurs there will be a vast body of data and expertise to find the best solution, particularly in relation to potential control measures and monitoring of microbial communities.
Surveillance of microbial communities can be used as a standard way to control and optimise wastewater treatment processes – it will be possible, if the identity and function of the most microorganisms is known, to determine whether the correct processes are in place, whereas today this is hard to resolve.
While the current focus is on bacteria, in principle it is possible to look at other biota and organisms of interest. It should, for example, be possible to begin looking into antibiotic resistance genes to determine whether they pose a real threat or not, and to investigate the type of resistant genes released in wastewater effluent and are potentially applied to farmland, adding unwanted bacteria.
The improved knowledge about the microbes and the option for online analysis should also help with a key current issue – the desire to recover resources such as N, P and energy – and to optimise plants. Today, this is a great challenge, but knowledge of microbes should help considerably.
Online analysis in full-scale systems is currently being tested and some plants will begin to show results within next few months. The key is showing that the approach can be applied. It will be around two years before the major wastewater treatment plants, whose efficient operation is of great concern, begin to use it; in five years it will be routine.
Change in the wastewater industry tends to take a long time, but if we can show online analysis is viable, it is a relatively cheap technology and with the database it will be easy to use and also fast – sampling and results can be obtained within a day, rather than weeks or months. The work can also be undertaken by less skilled workers rather than requiring expert input, so we believe it will be very attractive to wastewater treatment operators and consultants.
References
Andersen, M.H., McIlroy, S.J., Nierychlo, M., Nielsen, P.H., Albertsen, M., 2018. Genomic insights into Candidatus Amarolinea aalborgensis gen. nov., sp. nov., associated with settleability problems in wastewater treatment plants. Systematic and Applied Microbiology. https://doi.org/10.1016/j.syapm.2018.08.001
Dueholm, M.S., Andersen, K.S., Petriglieri, F., McIlroy, S.J., Nierychlo, M., Petersen, J.F., Kristensen, J.M., Yashiro, E., Karst, S.M., Albertsen, M., Nielsen, P.H., 2019. Comprehensive ecosystem-specific 16S rRNA gene databases with automated taxonomy assignment (AutoTax) provide species-level resolution in microbial ecology.
Jiang, C. M.H. Andersen, M.G. Peces, M. Nierychlo, M.S. Dueholm, E. Yashiro, K.S. Andersen, R.H. Kirkegaard, S. Kucheryavskiy, L. Hao, J. Høgh, A.A. Hansen, Per H. Nielsen (2019): Associations between species-level microbial communities and key variable revealed in anaerobic digesters at 22 Danish wastewater treatment plants during a six-years survey.
Karst, S.M., Dueholm, M.S., McIlroy, S.J., Kirkegaard, R.H., Nielsen, P.H., Albertsen, M., 2018. Retrieval of a million high-quality, full-length microbial 16S and 18S rRNA gene sequences without primer bias. Nature Biotechnology. https://doi.org/10.1038/nbt.4045
McIlroy, S.J., Kirkegaard, R.H., McIlroy, B., Nierychlo, M., Kristensen, J.M., Karst, S.M., Albertsen, M., Nielsen, P.H., 2017. MiDAS 2.0: an ecosystem-specific taxonomy and online database for the organisms of wastewater treatment systems expanded for anaerobic digester groups. Database 2017. https://doi.org/10.1093/database/bax016
Nierychlo, M., K.S. Andersen, Y. Xu, N. Green, M. Albertsen, M.S. Dueholm, P.H. Nielsen (2019): Species-level microbiome composition of activated sludge and anaerobic digesters – introducing MiDAS 3 ecosystem-specific reference database and taxonomy.
MiDAS – the Microbial Database for Activated Sludge
The MiDAS project is a collaborative scheme that has taken place since 2006 between more than 50 wastewater treatment plants, consultants and the Center for Microbial Communities, Aalborg University. It includes regular DNA and FISH analyses of the microbial community composition, in-depth studies of focus topics, and workshops and reports for plant operators. It also includes development and maintenance of the MiDAS field guide, and various MiDAS-curated taxonomies to promote a common language between researchers in the field. The newest is MiDAS 3.6, which is a comprehensive 16S RNA gene reference database based on high-quality sequences derived from activated sludge and anaerobic digester systems, and an improved taxonomy.
The MiDAS 3 database was the framework for an update of the MiDAS field guide – an online resource linking the identity of microorganisms in wastewater treatment systems to the available data on their functional importance. The new guide contains a complete list of genera (>1800) and species (>4000) found in activated sludge and anaerobic digesters. The identity of the microbes is linked to information on their function, where available. The website also enables classification of sequences using MiDAS 3 taxonomy directly online. Complex microbial communities define the function of biological wastewater treatment systems and understanding the role of individual microbes requires knowledge of their identity, to which known and putative functions can be assigned.